Cookie Policy: This site uses cookies to improve your experience. You can find out more about our use of cookies in our Privacy Policy. By continuing to browse this site you agree to our use of cookies.
Innaxon
LPS from E. coli EH100 (Ra) TLRpure Sterile Solution

| Product Details | |
|---|---|
| Synonyms | Lipopolysaccharide (LPS) from E. coli strain EH100 (Ra), R-type (rough/mutant) LPS |
| Product Type | Chemical |
| Properties | |
| Source/Host Chemicals | Isolated and purified from E. coli strain EH100. |
| Purity Chemicals | Ultrapure. No detectable DNA, RNA and protein traces. |
| Formulation | Liquid. Colourless clear aqueous solution. |
| Concentration | 1mg/ml stabilised in sterile, double-distilled water (ddWater), without any additives. |
| Biological Activity |
Optimal concentration is dependent upon cell type, species, desired activation and analysis: 0.05-1.0μg/ml. Does not activate any TLR other than TLR4 as tested up to 50μg/ml in relevant cellular systems (macrophages). |
| Declaration | Manufactured by Innaxon. |
| Other Product Data |
TLRpure™: • Qualified Purity & Activity • High potency TLR4-specific Ligands • Ultrapure (no detectable protein, RNA and DNA) • Tested on TLR4 KO murine macrophages • Standardised Aqueous Sterile Solutions • No purification or hazardous solubilisation • Excellent lot-to-lot consistency |
| Shipping and Handling | |
| Shipping | AMBIENT |
| Short Term Storage | +4°C |
| Long Term Storage | +4°C |
| Handling Advice |
Do not freeze. To yield a 100μg/ml (1,000-100x) stock solution add 100μl of LPS to 900μl endotoxin-free and sterile ddWater (Cat. No.: IAX-900-002) or 0.9% NaCl Solution (Cat. No.: IAX-900-003) and mix well. Prepare diluted LPS working solutions just prior to use, keep sterile. Ready-made solution is cell culture-grade. |
| Use/Stability | Stable for at least 2 years after receipt when stored at +4°C. |
| Documents | |
| MSDS |
 Download PDF Download PDF |
| Product Specification Sheet | |
| Datasheet |
 Download PDF Download PDF |
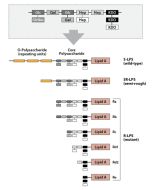
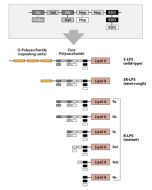
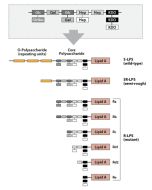
Activation of cells by LPS is mediated by the Toll-like receptor 4 (TLR4). For optimal interaction with LPS, TLR4 requires association with myeloid differentiation protein 2 (MD-2). According to current consensus activation of TLR4 is preceded by the transfer of LPS to membrane-bound (m) or soluble (s) CD14 by LPS-binding protein (LBP). Re-form LPS and lipid A, but not S-form LPS, are capable of inducing TNF-α responses also in the absence of CD14. LPS, synthesized by most wild-type (WT) Gram-negative bacteria (S-form LPS), consists of three regions, the O-polysaccharide chain, which is made up of repeating oligosaccharide units, the core oligosaccharide and the lipid A, which harbors the endotoxic activity of the entire molecule. R-form LPS synthesized by the so-called rough (R) mutants of Gram-negative bacteria lacks the O-specific chain. Furthermore, the core-oligosaccharide may be present in different degrees of completion, depending on the class (Ra to Re) to which the mutant belongs. LPS are amphipathic molecules whose hydrophobicity decreases with increasing length of the sugar part. Based upon these differences, S- and R-form LPS show marked differences in the kinetics of their blood clearance and cellular uptake as well as in the ability to induce oxidative burst in human granulocytes and to activate the host complement system.
- A new method for the extraction of R lipopolysaccharides: C. Galanos, et al.; Eur. J. Biochem. 9, 245 (1969)
- Electrodialysis of lipopolysaccharides and their conversion to uniform salt forms: C. Galanos & O. Lüderitz; Eur. J. Biochem. 54, 603 (1975)
- Immunoblot analysis of the R-form lipopolysaccharide from Salmonella S forms: S. Schlecht, et al.; Zentralbl. Bakteriol. 277, 288 (1992)
- Isolation and purification of R-form lipopolysaccharides: C. Galanos & O. Lüderitz; Methods in Carbohydrate Chem. 9, 11 (1993)
- The influence of low growth temperature on the amount of free R lipopolysaccharide, on the expression of R-core determinants and on O-chainlengths in Salmonella S forms: S. Schlecht & H. Mayer; Zentralbl. Bakteriol. 280, 448 (1994)
- Severe COVID-19 is marked by a dysregulated myeloid cell compartment: J. Schulte-Schrepping, et al.; Cell 182, 1419 (2020)
- Phenolic Compounds of Red Wine Aglianico del Vulture Modulate the Functional Activity of Macrophages via Inhibition of NF-κB and the Citrate Pathway: A. Santarsiero, et al.; Oxid. Med. Cell. Longev. 2021, 5533793 (2021)
- The Lipopolysaccharide Mutant Re-LPS Is a Useful Tool for Detecting LPS Contamination in Rheumatoid Synovial Cell Cultures: H. Kohono, et al.; Pathobiology (2021)