Cookie Policy: This site uses cookies to improve your experience. You can find out more about our use of cookies in our Privacy Policy. By continuing to browse this site you agree to our use of cookies.
AdipoGen Life Sciences
SARS-CoV-2 Spike Protein (D614G) (Stable Trimer) (rec.) (His)
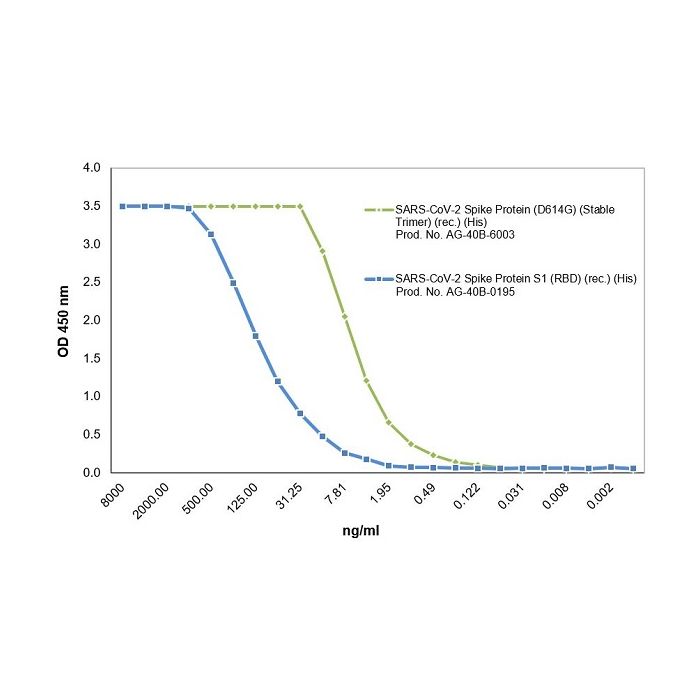
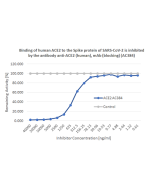
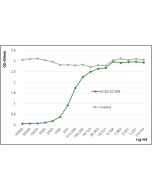
Method: SARS-CoV-2 Spike Protein (D614G) (Stable Trimer) (rec.) (His) (Prod. No. AG-40B-6003) or SARS-CoV-2 Spike Protein S1 (RBD) (rec.) (His) (Prod. No. AG-40B-0195) are coated at 50nM overnight at 4°C. ACE2 (human) (rec.) (Prod. No. AG-40B-0192) is added (starting at a concentration of 8000 ng/ml with a twofold serial dilution) during one hour at RT and the interaction is then detected using an anti-Flag (HRP).
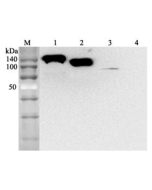
Method: SARS-CoV-2 Spike Protein (D614G) (Stable Trimer) (rec.) (His) has been injected in a SEC column ENrich SEC 650 10 x 300 mm (Bio-Rad).
| Product Details | |
|---|---|
| Synonyms | 2019-nCoV Spike Protein S |
| Product Type | Protein |
| Properties | |
| Source/Host | HEK 293 cells |
| Sequence |
Full-length soluble SARS-CoV-2 Spike protein (aa 16-1213) including a foldon trimerization motif, mutated Furin recognition site (R682S, R685S) and two stabilizing mutations (K986P and V987P) is fused at the C-terminus to a His-tag. |
| Crossreactivity | Human |
| Application |
Structural biology research / Vaccine development / Serological diagnostic kit development / Neutralizing antibody screening |
| Biological Activity |
Binds to the human SARS-CoV-2 receptor ACE2. Binds to anti-SARS-CoV-2 Spike antibodies in serum or plasma. |
| MW | ~190kDa (SDS-PAGE) |
| Purity | ≥90% (SDS-PAGE) |
| Endotoxin Content | <1EU/μg purified protein (LAL test). |
| Concentration | 0.2mg/ml after reconstitution. |
| Reconstitution | Reconstitute with 250µl endotoxin-free water. |
| Accession Number | P0DTC2 |
| Formulation | Lyophilized. Contains PBS and 0.5% trehalose. |
| Other Product Data |
UniProt link P0DTC2: SARS-CoV-2 S Protein |
| Shipping and Handling | |
| Shipping | BLUE ICE |
| Short Term Storage | +4°C |
| Long Term Storage | -20°C |
| Handling Advice |
After opening, prepare aliquots and store at -20°C. Avoid freeze/thaw cycles. Centrifuge lyophilized vial before opening and reconstitution. For maximum product recovery after thawing, centrifuge the vial before opening the cap. |
| Use/Stability | Stable for at least 6 months after receipt when stored at -20°C. |
| Documents | |
| MSDS |
 Download PDF Download PDF |
| Product Specification Sheet | |
| Datasheet |
 Download PDF Download PDF |
SARS-CoV-2 shares 79.5% sequence identity with SARS-CoV and is 96.2% identical at the genome level to the bat coronavirus BatCoV RaTG133, suggesting it had originated in bats. The original Wuhan strain of the virus has become quickly replaced by its more transmissible variant, mainly determined by a single amino acid point mutation D614G.
The coronaviral genome encodes four major structural proteins: the Spike (S) protein, Nucleocapsid (N) protein, Membrane/Matrix (M) protein and the Envelope (E) protein. The SARS Envelope (E) protein contains a short palindromic transmembrane helical hairpin that seems to deform lipid bilayers, which may explain its role in viral budding and virion envelope morphogenesis. The SARS Membrane/Matrix (M) protein is one of the major structural viral proteins. It is an integral membrane protein involved in the budding of the viral particles and interacts with SARS Spike (S) protein and the Nucleocapsid (N) protein. The N protein contains two domains, both of them bind the virus RNA genome via different mechanisms.
The CoV Spike (S) protein assembles as trimer and plays the most important role in viral attachment, fusion and entry. It is composed of a short intracellular tail, a transmembrane anchor and a large ectodomain that consists of a receptor binding S1 subunit (RBD domain) and a membrane-fusing S2 subunit. The S1 subunit contains a receptor binding domain (RBD), which binds to the cell surface receptor angiotensin-converting enzyme 2 (ACE2) present at the surface of epithelial cells. It has been demonstrated that certain mutations and the inclusion of trimerization motif can stabilize recombinant Spike protein trimers.
The recombinant protein SARS-CoV-2 Spike Protein (D614G) (Stable Trimer) (rec.) (His) could be useful for structural biology research, vaccine development, serological diagnostic kit development or neutralizing antibody screening.