Cookie Policy: This site uses cookies to improve your experience. You can find out more about our use of cookies in our Privacy Policy. By continuing to browse this site you agree to our use of cookies.
JaICA
Nuclease P1 (from Penicillium citrinum)
As low as
1891
CHF
CHF 1’891.00
In stock
Only %1 left
JAI-OPNUCP1500 UnitsCHF 1’891.00

| Product Details | |
|---|---|
| Product Type | Protein |
| Properties | |
| Sequence | A zinc dependent glycoprotein consisting of 270 amino acid residues. |
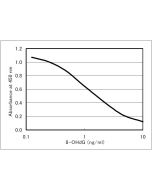
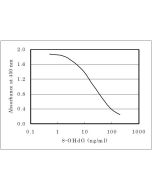
| Application | Used for DNA samples pretreatment to detect 8-OHdG. |
| Biological Activity | 3′-5′-Phosphodiesterase: One unit will liberate ~1.0μmol of acid soluble nucleotides from RNA per min at pH 5.3 at 37°C. 3′-Nucleotidase: One unit will hydrolyze ~4.0μmol of orthophosphate from 3′-AMP per min at pH 7.2 at 37°C. |
| MW | 42-50 kDa |
| Formulation | Lyophilized. |
| Other Product Data |
The enzyme has an optimal temperature of approximately 70 °C, but for a long incubation, a temperature below 60 °C is more suitable. It is stable in the pH range of 5 - 8. Optimal pH: 5.3 (RNA and heat-denatured DNA); 7.2 (3'-AMP) Heat Stability: Stable below 60°C at pH 5.3 for 30 minutes. |
| Declaration | Manufactured by JaICA. |
| Shipping and Handling | |
| Shipping | BLUE ICE |
| Short Term Storage | +4°C |
| Long Term Storage | +4°C |
| Use/Stability | Stable for at least 1 year after receipt when stored at +4°C. |
| Documents | |
| Product Specification Sheet | |
| Datasheet |
 Download PDF Download PDF |
Description
Catalyzes the nonspecific endonucleolytic cleavage of single stranded DNA and RNA to yield nucleoside 5′-phosphates and 5′-phospho-oligonucleotides. Nuclease P1 hydrolyzes both 3'-5'-phosphodiester bonds in RNA and heat-denatured DNA and 3'-phosphomonoester bonds in mono- and oligonucleotides terminated by 3'-phosphate without base specificity. The enzyme does not attack double-stranded nucleic acids, especially in the presence of more than 400mmol/l of sodium chloride at pH 6.0.
Product References
- The interactions of dopamine and oxidative damage in the striatum of patients with neurodegenerative diseases: H. Li, et al.; J. Neurochem. 152, 235 (2020)